 
There are a number of 5′ and 3′ modifications that can be selected, and will change the molecular weight and in some cases the absorbance coefficient for the oligonucleotide. The composition of the oligonucleotide (DNA or RNA, single-stranded or double-stranded) can be selected and will change many of the calculations, although the absorption coefficients are only accurate for single-stranded oligonucleotides. The nanomolar concentration of primer in the hybridization solution can also be entered and will adjust the nearest neighbor melting temperature. The millimolar concentration of salt can be entered and will adjust the salt adjusted and nearest neighbor melting temperature calculations. 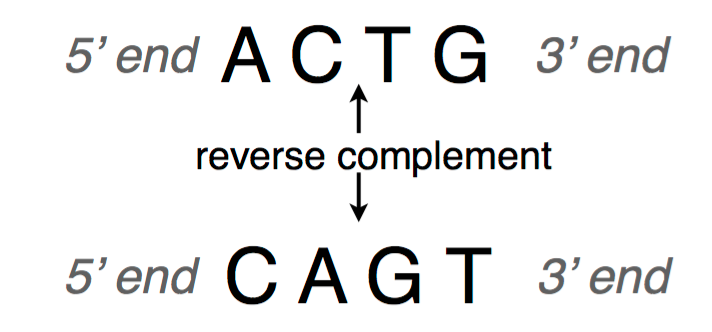
The absorbance at 260 nm (A260) can be entered for the oligonucleotide, and will result in the calculation of the micromolar concentration of oligonucleotide as well as the micrograms of the oligonucleotide present in a 1 ml solution with that absorbance. Once the user has entered a sequence, several additional options can be selected or entered. Other options include the swapping of the entered sequence for its complement, submitting the sequence to the NCBI BLAST site, calculating self-complementarity between two identical oligonucleotide molecules, and calculating potential intra-molecular hairpin loop formation.Įntry and main calculation screen for OligoCalc. The user can also select options such as single- or double-stranded DNA and RNA molecules (ssDNA is the default), and the user can select from more than seventy 5′ and 3′ chemical modifications that refine the molecular weight and absorbance calculations for oligonucleotides with those modifications. In addition to these calculations, the user can enter absorbance readings to calculate the concentration of the oligonucleotide in ng/μmol and micrograms present in 1 ml of solution, and enter the predicted salt and/or primer concentrations in the final hybridization solution to more accurately predict the melting temperature. Performing these calculations are all straightforward-enter the nucleotide sequence into a textbox, and hit return or click on the ‘Calculate’ button. 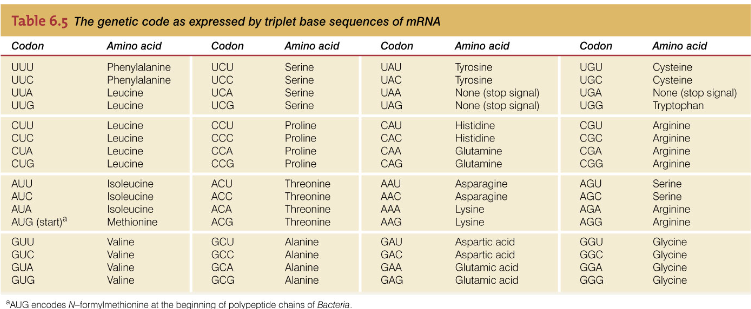
OligoCalc provides the results of three common melting temperature calculations. The recent interest and availability of biological applications for siRNAs has resulted in the addition of RNA oligonucleotides as common laboratory reagents, and OligoCalc can be used to calculate the properties of single-stranded and double-stranded RNA as well as DNA. OligoCalc provides a convenient web interface for calculating the physical properties of DNA and RNA oligonucleotides including melting temperature, molecular weight, %GC content and absorbance coefficient for a given oligonucleotide sequence. With the availability of completely sequenced genomes, various genomic and array technologies including DNA microarrays and bead arrays have made oligonucleotides even more important reagents. Even prior to PCR, DNA oligonucleotides were used extensively in molecular biology as primers and as probes. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
